library(tidyverse)Warning: package 'ggplot2' was built under R version 4.4.3Warning: package 'tibble' was built under R version 4.4.3Warning: package 'tidyr' was built under R version 4.4.3Warning: package 'readr' was built under R version 4.4.3Warning: package 'purrr' was built under R version 4.4.3Warning: package 'dplyr' was built under R version 4.4.3Warning: package 'lubridate' was built under R version 4.4.3── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.2.0 ✔ readr 2.2.0
✔ forcats 1.0.1 ✔ stringr 1.6.0
✔ ggplot2 4.0.2 ✔ tibble 3.3.1
✔ lubridate 1.9.5 ✔ tidyr 1.3.2
✔ purrr 1.2.1
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary(here)here() starts at /Users/ayllaermland/Downloads/BIOS8060E/AyllaErmland-portfolio# Load dataset
data <- read_csv(here("fitting-exercise", "Mavoglurant_A2121_nmpk.csv"))Rows: 2678 Columns: 17
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
dbl (17): ID, CMT, EVID, EVI2, MDV, DV, LNDV, AMT, TIME, DOSE, OCC, RATE, AG...
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.# Inspect structure
glimpse(data)Rows: 2,678
Columns: 17
$ ID <dbl> 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, …
$ CMT <dbl> 1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 1, 2, 2, 2, 2, 2,…
$ EVID <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0,…
$ EVI2 <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0,…
$ MDV <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0,…
$ DV <dbl> 0.00, 491.00, 605.00, 556.00, 310.00, 237.00, 147.00, 101.00, 72.…
$ LNDV <dbl> 0.000, 6.196, 6.405, 6.321, 5.737, 5.468, 4.990, 4.615, 4.282, 3.…
$ AMT <dbl> 25, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 25, 0, 0, 0, 0, …
$ TIME <dbl> 0.000, 0.200, 0.250, 0.367, 0.533, 0.700, 1.200, 2.200, 3.200, 4.…
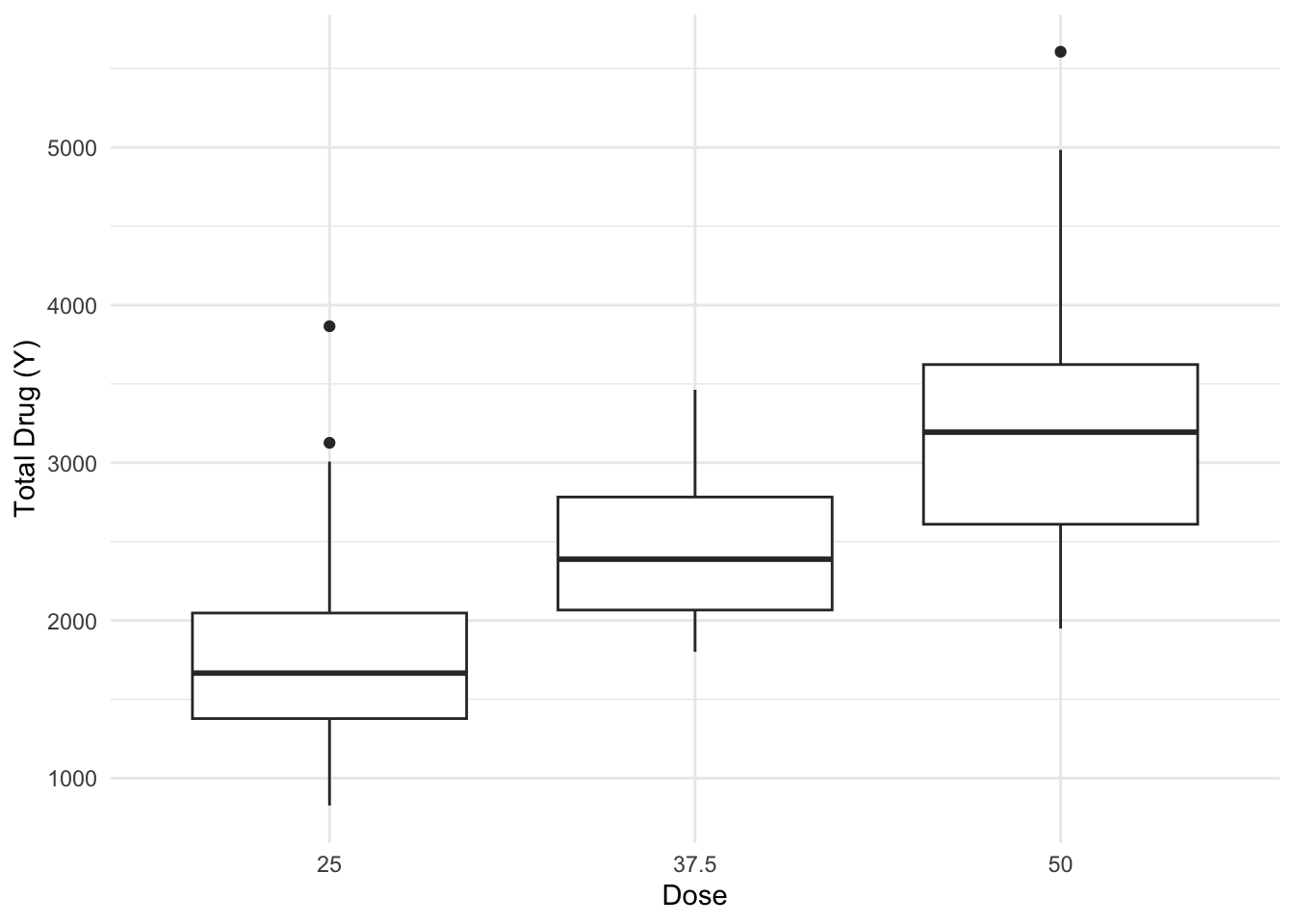
$ DOSE <dbl> 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 2…
$ OCC <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,…
$ RATE <dbl> 75, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 150, 0, 0, 0, 0,…
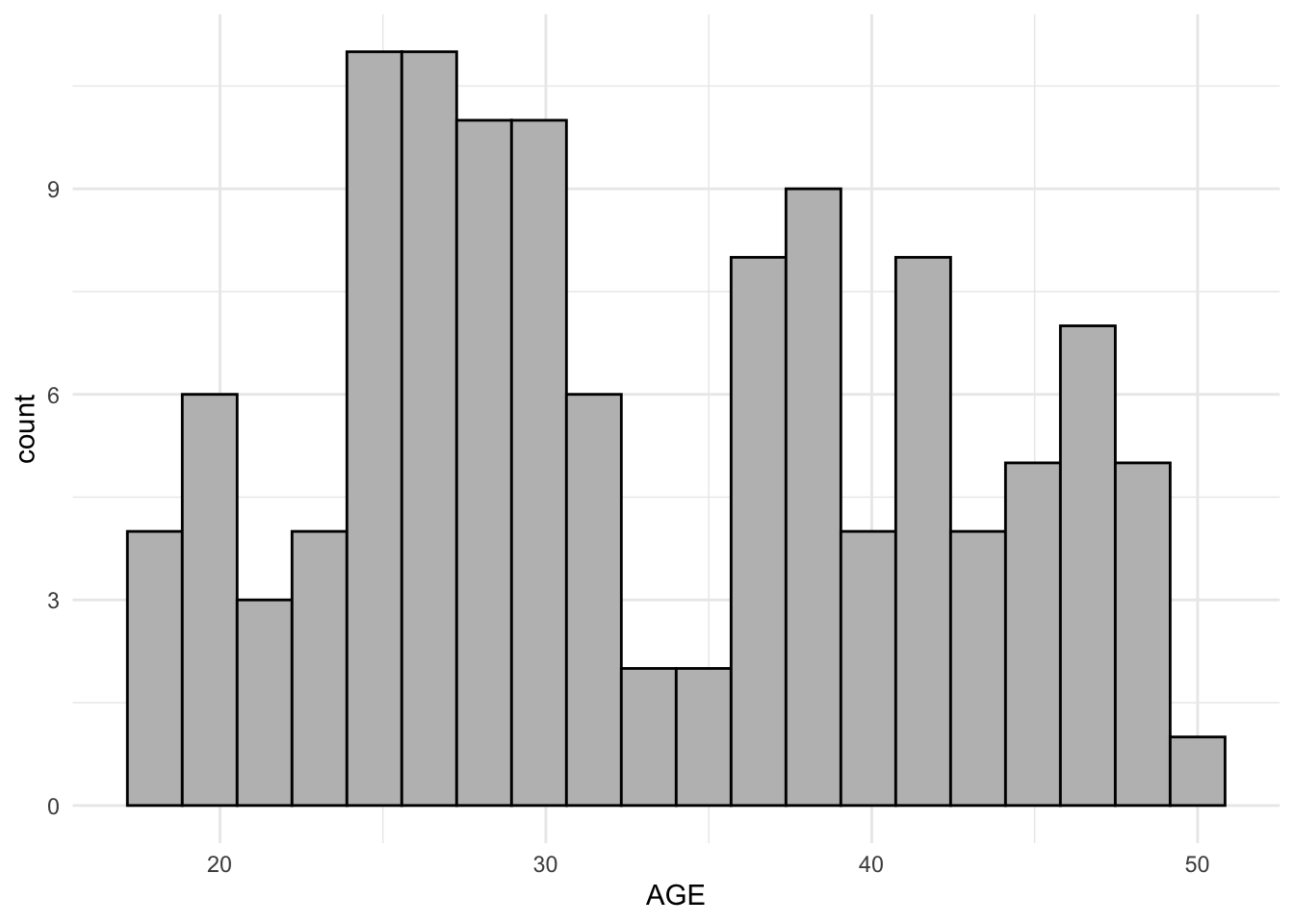
$ AGE <dbl> 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 2…
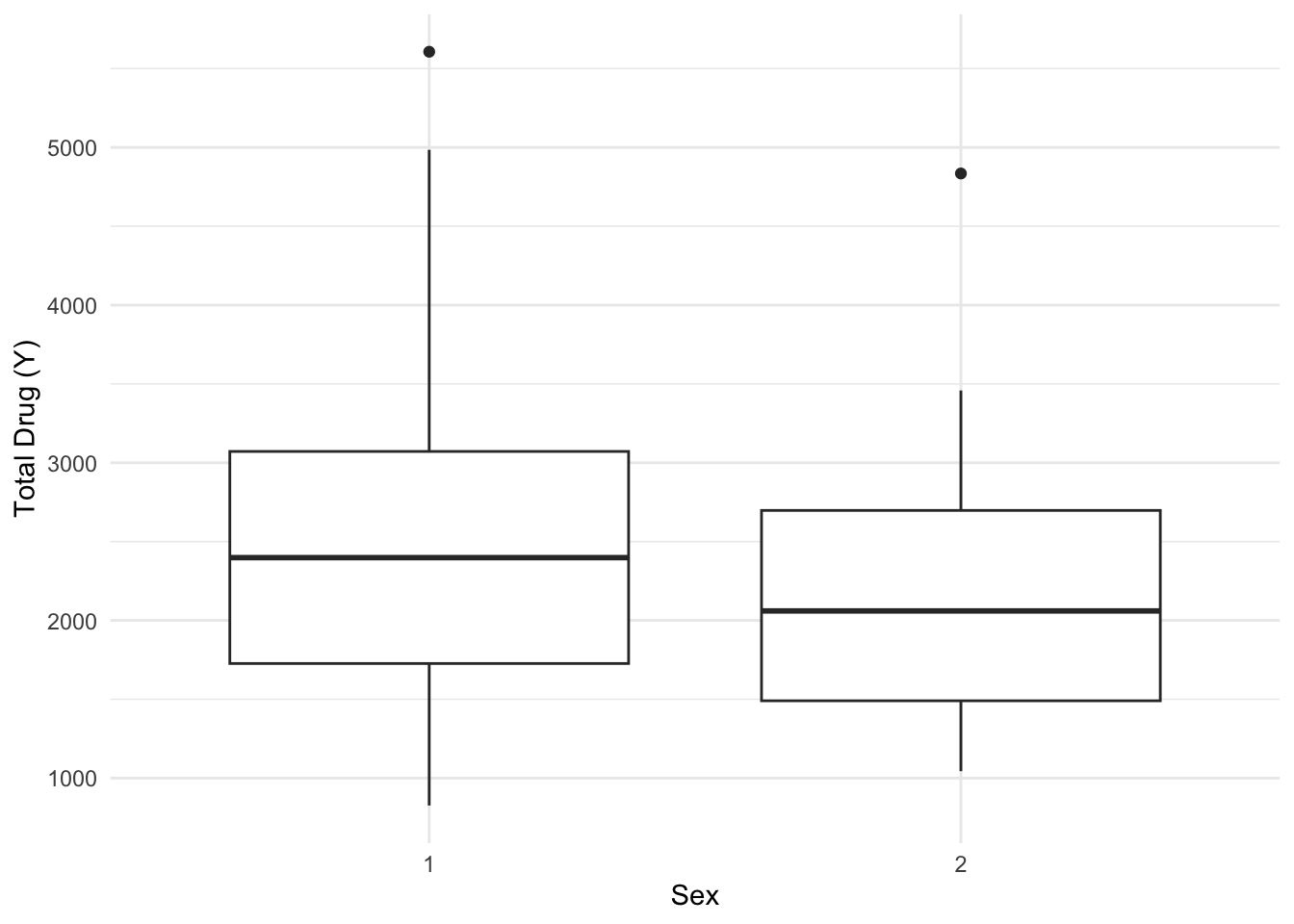
$ SEX <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,…
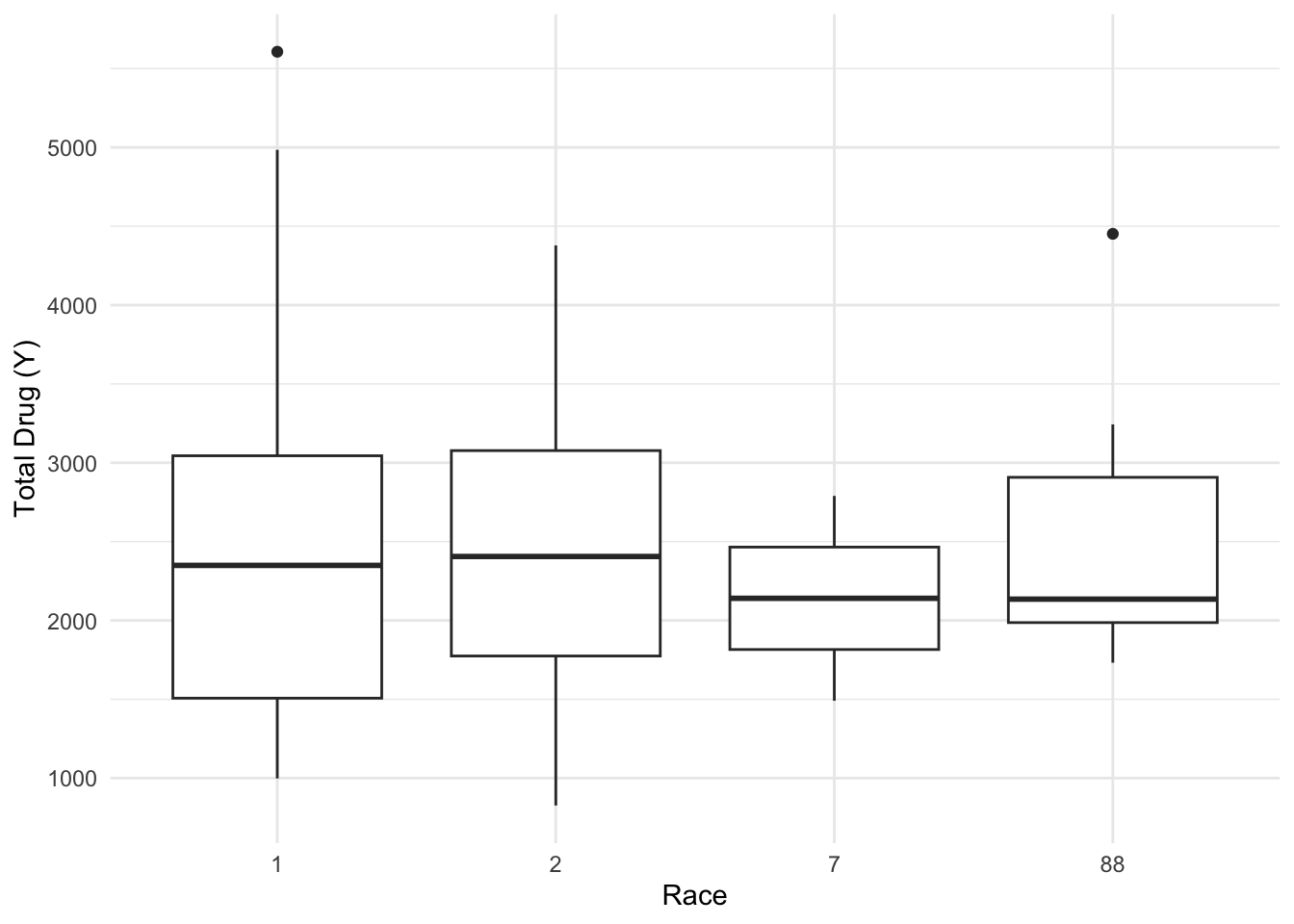
$ RACE <dbl> 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2,…
$ WT <dbl> 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3,…
$ HT <dbl> 1.769997, 1.769997, 1.769997, 1.769997, 1.769997, 1.769997, 1.769…str(data)spc_tbl_ [2,678 × 17] (S3: spec_tbl_df/tbl_df/tbl/data.frame)
$ ID : num [1:2678] 793 793 793 793 793 793 793 793 793 793 ...
$ CMT : num [1:2678] 1 2 2 2 2 2 2 2 2 2 ...
$ EVID: num [1:2678] 1 0 0 0 0 0 0 0 0 0 ...
$ EVI2: num [1:2678] 1 0 0 0 0 0 0 0 0 0 ...
$ MDV : num [1:2678] 1 0 0 0 0 0 0 0 0 0 ...
$ DV : num [1:2678] 0 491 605 556 310 237 147 101 72.4 52.6 ...
$ LNDV: num [1:2678] 0 6.2 6.41 6.32 5.74 ...
$ AMT : num [1:2678] 25 0 0 0 0 0 0 0 0 0 ...
$ TIME: num [1:2678] 0 0.2 0.25 0.367 0.533 0.7 1.2 2.2 3.2 4.2 ...
$ DOSE: num [1:2678] 25 25 25 25 25 25 25 25 25 25 ...
$ OCC : num [1:2678] 1 1 1 1 1 1 1 1 1 1 ...
$ RATE: num [1:2678] 75 0 0 0 0 0 0 0 0 0 ...
$ AGE : num [1:2678] 42 42 42 42 42 42 42 42 42 42 ...
$ SEX : num [1:2678] 1 1 1 1 1 1 1 1 1 1 ...
$ RACE: num [1:2678] 2 2 2 2 2 2 2 2 2 2 ...
$ WT : num [1:2678] 94.3 94.3 94.3 94.3 94.3 94.3 94.3 94.3 94.3 94.3 ...
$ HT : num [1:2678] 1.77 1.77 1.77 1.77 1.77 ...
- attr(*, "spec")=
.. cols(
.. ID = col_double(),
.. CMT = col_double(),
.. EVID = col_double(),
.. EVI2 = col_double(),
.. MDV = col_double(),
.. DV = col_double(),
.. LNDV = col_double(),
.. AMT = col_double(),
.. TIME = col_double(),
.. DOSE = col_double(),
.. OCC = col_double(),
.. RATE = col_double(),
.. AGE = col_double(),
.. SEX = col_double(),
.. RACE = col_double(),
.. WT = col_double(),
.. HT = col_double()
.. )
- attr(*, "problems")=<externalptr> head(data)# A tibble: 6 × 17
ID CMT EVID EVI2 MDV DV LNDV AMT TIME DOSE OCC RATE AGE
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 793 1 1 1 1 0 0 25 0 25 1 75 42
2 793 2 0 0 0 491 6.20 0 0.2 25 1 0 42
3 793 2 0 0 0 605 6.40 0 0.25 25 1 0 42
4 793 2 0 0 0 556 6.32 0 0.367 25 1 0 42
5 793 2 0 0 0 310 5.74 0 0.533 25 1 0 42
6 793 2 0 0 0 237 5.47 0 0.7 25 1 0 42
# ℹ 4 more variables: SEX <dbl>, RACE <dbl>, WT <dbl>, HT <dbl>